A pseudogene is a piece of DNA that looks like a gene but
doesn't make a working protein. It is the molecular
equivalent of a photocopy of a key — recognisable, the right shape,
but somewhere along the way the cuts didn't quite line up. Most are
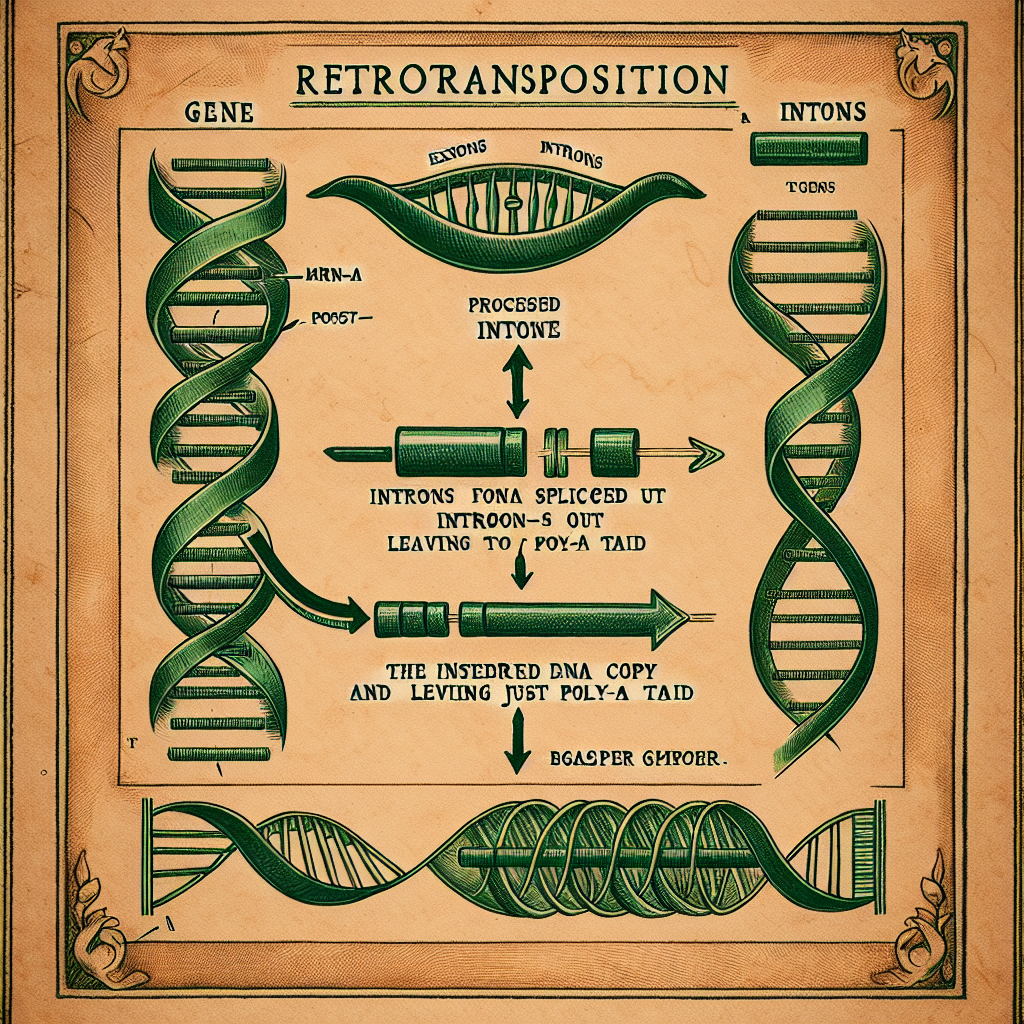
relics of duplications that happened tens of millions of years ago,
still sitting in our genome long after they stopped doing what their
ancestors did.
For decades, biologists treated them as evolutionary debris. The term
itself — coined by Jacq, Miller and Brownlee in 1977 — was slightly
dismissive: pseudo, "false." A century of textbooks pushed
the same line: pseudogenes are mistakes, gene fossils, the parts of
the genome that don't matter.
Then, around 2010, a series of papers showed that some pseudogenes
produce RNAs that regulate the expression of their parent
genes. One of them, PTENP1, turned out to be a
tumour suppressor in its own right. The clean line between "gene"
and "junk" stopped being clean.
Some pseudogenes do nothing. Some regulate cancer. Some are diagnostic
landmines. We mostly don't yet know which is which.